Brogi Francesco

OC23 – Brogi Francesco
Department of Pharmaceutical Sciences, University of Perugia, Italy
francesco.brogi@dottorandi.unipg.it
| Chemical Probes for Mono-ADP-Ribosylating enzymes (mono-ARTs) inhibition and degradation |
| Brogi Francesco1, Juho Alaviuhkola2,Lari Lehtiӧ2, Tabarrini Oriana 1 1 Department of Pharmaceutical Sciences, University of Perugia, Italy 2 Faculty of Biochemistry and Molecular Medicine, Biocenter Oulu, University of Oulu, Finland |
| Abstract Human ADP-ribosyltransferases (ARTs), PARP enzymes and tankyrases, form a family of 17 proteins that catalyze ADP-ribosylation, a post-translational modification of proteins that regulates numerous cellular processes[1]. Poly-ARTs(PARP1-2) are well characterised, and six PARP1 inhibitors are in clinical use as anticancer agents, with a mechanism of action based on the synthetic lethality principle. The first of these inhibitors, Olaparib was approved in 2014, representing a success in the field of the precision medicine[2]. In contrast, mono-ARTs (PARP3–17), the discovery of which began in 2010 with PARP10, remain largely unexplored despite growing evidence of their involvement in oncological, immune, infectious and neurodegenerative diseases.[3]. This project aims to develop selective PARPs inhibitors as chemical probes to elucidate their biology and validate them as innovative therapeutic targets. Starting from OUL232 (figure 1), the most potent and selective PARP10 inhibitor reported to date[3], we have designed and synthesized a large series of [1,2,4] triazolo[3,4-b]benzothiazole derivatives in an attempt to achieve selectivity especially towards PARP7, PARP11 or PARP14, which are potential immuno-oncology targets. In particular, PARP14 controls inflammatory pathways and interferon responses, PARP7 modulates interferon signaling and nuclear receptors such as estrogen and androgen receptors, while PARP11 regulates interferon signaling and promotes immunosuppressive tumor environments[1]. In parallel, we are also preparing PROteolysis TArgeting Chimeras (PROTAC) and Hydrophobic Tagging (HyT) derivatives, to investigate whether PARP degradation could provide therapeutic advantages. The presentation will cover the design and synthesis of PARP inhibitors and protein degraders, PARP inhibition and crystallographic studies. 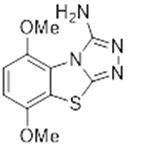 Figure 1. Structure of PARP10 inhibitor OUL232 |
| References: [1] Di Paola, S., Grimaldi, G. & Corda, D. Mono-ADP-ribosylating PARP enzymes in cellular signaling and disease. Journal of Cell Science, 2025, vol. 138 (14), https://doi.org/10.1242/jcs.263963. [2] Wang, Y. et al. The Potential of PARP Inhibitors as Antitumor Drugs and the Perspective of Molecular Design. J. Med. Chem. 2025, 68, 18–48, https://doi.org/10.1021/acs.jmedchem.4c02642 [3] Murthy, S. et al. [1,2,4]Triazolo[3,4-b]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes. J. Med. Chem. 2023, 66, 1301–1320, https://doi.org/10.1021/acs.jmedchem.2c01460; b) Compounds for use in the treatment of cancer and inflammatory conditions L Lehtiö, M Maksimainen, SN Murthy, O Tabarrini, MG Nizi, S Massari US Patent App. 18/834,003 |