Anouk Van Hauwermeiren
| Macrocyclization as a strategy for optimizing protacs targeting Aurora Kinase A |
| Anouk Van Hauwermeiren1, Simon Krols1, Fien Martens2, Hanne Van De Putte1, Muhammad Rishifi2, Kaat Durinck2, Serge Van Calenbergh1 1 Laboratory for Medicinal Chemistry, Faculty of Pharmaceutical Sciences, Ghent University, Belgium; 2 Department of Biomolecular Medicine, Faculty of Medicine & Health Sciences, Ghent University, Belgium |
| Abstract Previous efforts at the Laboratory for Medicinal Chemistry led to the discovery of proteolysis-targeting chimeras (PROTACs) directed against Aurora kinase A (AURKA). Although these demonstrated potent in vitro degradation activity in neuroblastoma cell lines, their in vivo efficacy was limited, primarily due to poor metabolic stability, which hindered further development. [1] To address this limitation, macrocyclization was explored as a structural optimization strategy. By restricting conformational flexibility and limiting enzymatic accessibility, macrocyclization may improve the potency, selectivity, and metabolic stability of small molecules. [2–3] Guided by the co-crystal structure of the selective AURKA inhibitor MK-5108 bound to its target (figure 1), a series of macrocyclic AURKA ligands was designed, synthesized, and elaborated into AURKA-targeting PROTACs. Biological evaluation demonstrated that these macrocyclic PROTACs retained degradation potency comparable to their non-macrocyclic counterparts, indicating that macrocyclization is well tolerated within the PROTAC context. Notably, the macrocyclic PROTACs exhibited significantly improved metabolic stability, particularly with regard to cytochrome P450-mediated metabolism. Collectively, these findings highlight macrocyclization as a promising strategy to optimize PROTACs for improved pharmacokinetic properties. 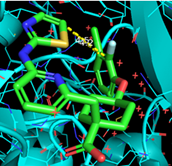 Figure 1. Co-crystal structure of MK-5108 bound to AURKA |
| References [1] Krols S, Rishfi M, et al. Second-Generation AURKA-Targeting PROTACs: Structural Optimization toward in Vivo Degradation in Neuroblastoma. J Med Chem. 2025 Nov 27;68(22):23962–76. doi:10.1021/acs.jmedchem.5c01271 [2] Driggers EM, Hale SP, et al. The exploration of macrocycles for drug discovery — an underexploited structural class. Nat Rev Drug Discov. 2008 Jul;7(7):608–24. doi:10.1038/nrd2590 [3] Amrhein JA, Knapp S, et al. Synthetic Opportunities and Challenges for Macrocyclic Kinase Inhibitors. J Med Chem. 2021 Jun 24;64(12):7991–8009. doi:10.1021/acs.jmedchem.1c00217 |
